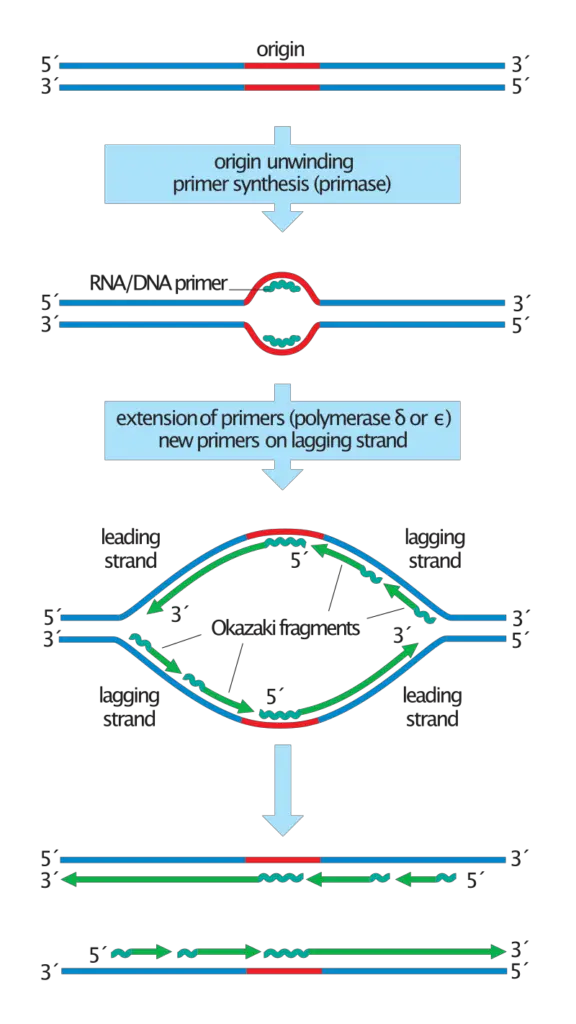
Selsing E, Wells RD, Alden CJ, Arnott S.Two contiguous conformations in a nucleic acid duplex. Selsing E, Wells RD, Early TA, Kearns DR.Structural analysis of spermine and magnesium ion binding to yeast phenylalanine transfer RNA. Molecular structure of r(GCG)d(TATACGC): a DNA-RNA hybrid helix joined to double helical DNA. Wang AH, Fujii S, van Boom JH, van der Marel GA, van Boeckel SA, Rich A.Cryocrystallography of biological macromolecules: a generally applicable method. Chemical synthesis of biologically active oligoribonucleotides using beta-cyanoethyl protected ribonucleoside phosphoramidites. RNA-linked nascent DNA fragments in Escherichia coli. A possible role for RNA polymerase in the initiation of M13 DNA synthesis. Nucleoside triphosphate termini in RNA polymerase products. Possible discontinuity and unusual secondary structure of newly synthesized chains. Okazaki R, Okazaki T, Sakabe K, Sugimoto K, Sugino A.Mechanism of DNA replication possible discontinuity of DNA chain growth. Okazaki R, Okazaki T, Sakabe K, Sugimoto K.Origin and direction of simian virus 40 deoxyribonucleic acid replication. On the mechanism of DNA replication in mammalian chromosomes. THE REPLICATION OF DNA IN ESCHERICHIA COLI. Links to PubMed are also available for Selected References. Get a printable copy (PDF file) of the complete article (1.4M), or click on a page image below to browse page by page. Full textįull text is available as a scanned copy of the original print version. The RNA trimer may, therefore, lock the complete fragment in an A-type conformation. Although the base-pair geometry, particularly in the central TATA part, is distorted, there is no evidence for a transition from the A- to the B-type conformation at the junction between RNA.DNA hybrid and DNA duplex. The fragment adopts an overall A-type conformation with 11 residues per turn. The DNA oligonucleotide d(GGGTATACGC) and the chimeric RNA-DNA oligonucleotide r(GCG)d(TATACCC) were combined to form a synthetic Okazaki fragment and its three-dimensional structure was determined by x-ray crystallography. They are composed of the growing DNA strand primed by RNA and the template strand. Okazaki fragments are a part of the lagging strand, and lagging strands give space to Okazaki fragments which is responsible for the continuous growth of Okazaki fragments.In DNA replication, Okazaki fragments are formed as double-stranded intermediates during synthesis of the lagging strand.While lagging strands continue formation with the short RNA sequences and maintain the development of Okazaki fragments. Okazaki fragments are formed by the action undertaken by the strand.Okazaki fragments are 100 to 200 nucleotides long, but the average size of lagging strands is 3000 nucleotides long.Okazaki fragments are comparatively shorter in size, but lagging strands, when compared, are much longer in size.It is responsible for the continuous growth of Okazaki fragments. Okazaki Fragments are small sections of DNA that are dependent on lagging strands to be produced, whereas lagging strands are nothing but replicated DNA strands.Main Differences Between Okazaki Fragments and Lagging Strand Without the work of lagging strands, the process of making Okazaki fragments will get disrupted, and ultimately the proper DNA formation will be compromised. DNA ligase is another requirement in the joining of Okazaki fragments. The formation of Okazaki fragments requires a new RNA primer every time. 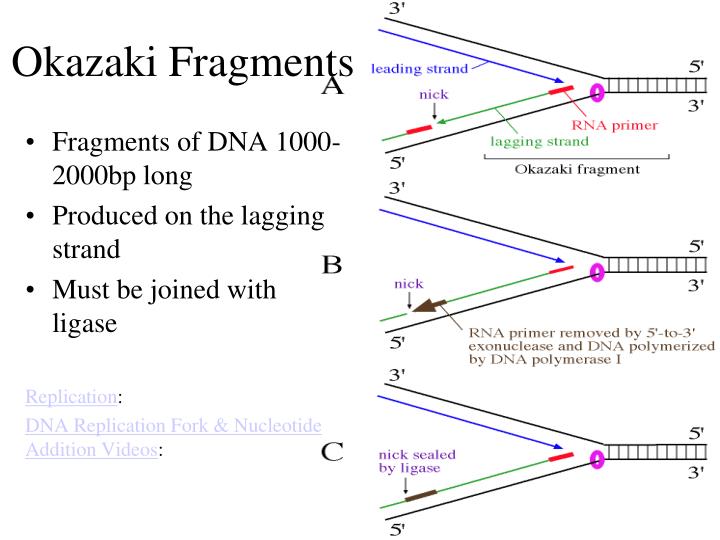
When lagging strands start to synthesize, they produce Okazaki fragments. The formation of lagging strands is a slow process. It is capable of opening up in the 5′ to 3′ direction and grows in the 3′ to 5′ direction. The growth of lagging strands takes place when it discontinuously constructs Okazaki Fragments. Japanese biologists Tsuneko Okazaki discovered them in the 1960s with the help of Reiji and other colleagues. These fragments are named after their founder. These are short, roughly we can measure their average size to be 150 to 200 nucleotides. Okazaki fragments are cycles of DNA nucleotides. Lagging strands give space to Okazaki fragments. Its average size is 3000 nucleotides long. Lagging strands continue formation with the short RNA sequences and maintain the development of Okazaki fragments. It is formed by the action undertaken by the strand. These are small sections of DNA that are produced by the uninterrupted process of lagging strands. Comparison Table Parameters of Comparison These are large in size, and it is the other half of the leading strand which grows without a gap, unlike lagging strands. The formation of Okazaki fragments is not possible unless lagging strands perform their duties.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
